Cases Collection
A total of 302 patients who received orthodontic treatment at the Department of Orthodontics, West China Hospital of Stomatology in Chengdu, China, from 2014 to 2018 were included in this study. The inclusion criteria were fixed labial appliance patients with full permanent dentition (except for second or third molars) without functional appliance treatment or orthognathic surgery. Their medical records before orthodontic treatment were collected, including demographic information, extraoral photos, intraoral photos, pretreatment dental casts and lateral cephalometric measurements6,29. Twenty-four commonly used feature variables were extracted from these clinical records as input features. The input features were preprocessed to ensure that all of them were quantified before being used for model training. Nonquantitative data were converted into numerical values by the encoding method. Supplementary Table S3 shows the detailed features used in the ANNs and how the nonquantitative data were encoded. All treatment planning was carefully performed by Dr. Zhao and Dr. Tang, who are both orthodontic specialists and have 26 and 12 years of clinical work experience, respectively. This study was approved by the West China Hospital of Stomatology Institutional Review Board (WCHSIRB-D-2018-094). Informed consent was obtained from all participants or their legal guardians. Informed consent for publication of the medical records of four example patients in an online open-access publication was also obtained. All experiments were performed in accordance with relevant guidelines and regulations.
The composition of the cases and datasets
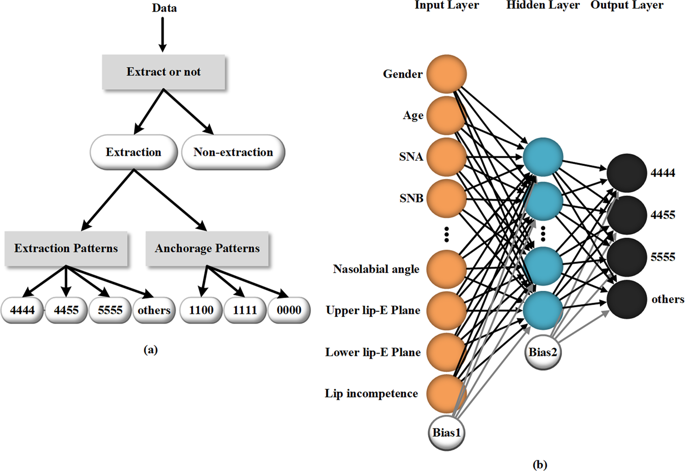
Among the total population, 222 persons were extraction cases, accounting for 73.5%, and the other 80 persons were nonextraction cases, accounting for 26.5%. The tooth extraction patterns were divided into four types: maxillary and mandibular first premolar extraction (4444), maxillary first premolar and mandibular second premolar extraction (4455), maxillary and mandibular second premolar extraction (5555) and other extraction patterns including only maxillary first premolar extraction, maxillary second premolar and mandibular first premolar extraction. These four patterns comprise 41.9%, 19.8%, 18.5% and 18.5% of the extraction cases, respectively. The anchorage patterns included three types, i.e., maxillary maximum anchorage (1100), maxillary and mandibular maximum anchorage (1111), and no use of maximum anchorage (0000), accounting for 29.7%, 21.6% and 48.6% of the extraction cases, respectively. Descriptions of the extraction patterns and anchorage patterns are shown in Table 3.
The dataset is split into a training set, a validation set and a test set. The neural networks do not have access to the test set during the training process until the final evaluation of the accuracy. The reserve part of the dataset is split into a training set and a validation set with a ratio of 3/1, which is optimized according to the learning curve30. The training set is used to update the weights of the network. The validation set is used to avoid overfitting13. Considering that we had a smaller dataset, we used a greater percentage of data to test the models. Therefore, the training set, validation set and test set were set with a typical 60/20/20 split to maintain a balance between the sets. Cases with different tags were randomly distributed to the three datasets in each simulation so that the proportions of various cases are similar among the three sets, reducing the additional bias introduced by the data partitioning process. There are 222 extraction cases; thus, the 222 cases are used in the neural network models for predicting extraction patterns and anchorage patterns. The number and percentage of different kinds of treatment plans in each set are shown in Table 4.
Network models
All three neural networks used in this work are three-layer MLPs. Each MLP consists of three full connection layers. The MLP used to determine extraction patterns is illustrated in Fig. 1b. The activation function of the hidden layer is tanh. A softmax layer of 4 outputs is applied at the end of the model31,32. The cross-entropy33,34CEtanh is given by
$$CE=-,tast mathrm{log}(y)-(1-t)ast mathrm{log}(1-y)$$
(1)
where t is the target value and y is the output of the MLP. Equation (1) returns a numerical value approaching infinity, which heavily penalizes output when y approaches −1 or 1. CEtanh approaches its minimum value when y approaches t. The weight and bias values are updated according to the scaled conjugate gradient method35. Although minimizing CEtanh leads to a good accuracy of classification, considerably minimizing CEtanh may cause overfitting. The dropout method is used to prevent overfitting36,37. The detailed training setting including learning rate, number of epochs, batch size, et al., are provided in Supplementary Note 4.
For the extraction prediction, the model outputs a probability of extraction. We define a determination of extraction treatment for each case as the probability of extraction being higher than a cutoff value. The algorithm computes sensitivity and specificity by testing a variety of cutoff values. Varying the cutoff point in the interval 0–1 generates a conventional ROC curve. Youden’s index38 is applied to obtain the optimum cutoff. If the probability is higher than the optimum cutoff, the case will be passed to the prediction of extraction patterns and anchorage patterns.
Relative contribution calculation of features and complement of the missing data
The PaD method, which is supposed to be the most useful method in giving the relative contribution and the contribution profile of the input factors, was used to evaluate the relative contribution calculation of the input features. The PaD method computes the partial derivatives of the ANN’s output with respect to the input to obtain the profile of the variations of the output for small changes of one input variable. For a network with ni inputs (where i represents the feature index and i = 1, 2, …, 24 in this work), one hidden tanh layer with nh neurons, and no outputs, the partial derivatives of the output yj with respect to input xj (where j represents the case index and j = 1, 2, …, 302 in this work) are:
$${d}_{ij}={S}_{j}sum _{h=1}^{{n}_{h}}{w}_{ho}(1-{I}_{hj}^{2}){w}_{ih}$$
(2)
where Sj is the derivative of the output neuron with respect to the input, which is the weights between the output neuron and hth hidden neuron, Ihj is the output of the hth hidden neuron, and wih is the weights between the ith input neuron and the hth hidden neuron.
Then, the relative contribution of the ANN’s output to the dataset with respect to the ith input feature can be calculated by a sum of the square partial derivatives as:
$$SS{D}_{i}=sum _{j=1}^{N}{d}_{ij}^{2}$$
(3)
where N is the data size and equals 302 in this work. The SSD values enable direct access to the influence of each input variable on the output.
We use the average value method, frequent value method, specific value method, median value method, and k-NN method to study their complement effects. The four traditional methods complement the missing data with the average value, the frequent value, the specified value (standard value of normal population), and the median value. The k-NN method is a method to look for the new case’s nearest neighbors from the complete cases and use an estimated value to replace the missing data26,39. This value is the weighted average of the values of its k nearest neighbors. We used 2-k-NN (2 nearest neighbors) and 3-k-NN (3 nearest neighbors) in this study. Each neighbor is given a weight of 1/d, where d is the distance to the neighbor. The neighbors are taken from the dataset for which the object property value is known.